So I’m an electron microscopy facility manager at an R1 university and have designed and implemented a number storage and data movement solutions, mostly for cryo-EM.
For $100k you should easily be able to hit 300 TB storage. In fact, you can probably hit about a petabyte for that budget. At the moment I can’t remember the exact chassis, but Supermicro sells a 4U with 36 bays. We have a single zpool in RAIDz 6 (or whatever the ZFS RAID 6 is, I can’t recall at the moment) on a system with three 12 drive vdevs, each with 16 TB drives, for about 480 TB. We started off with just a single vdev of 12 disks but you have to choose between scale-up vs scale-out at the beginning, and scale-out is complex and expensive, especially at first.
If you must have scale-out, going with something like Ceph is a pain in the ass and probably over-complex for your data needs, but something like MooseFS is simpler, and you can get off the ground faster while still scaling.
I know it’s in use at Purdue in their structural biology department, which is one of the premier ones in the world and generates an ungodly amount of data, and they are pretty happy with it. When I talked to them they said they were just adding a new off-the-shelf SuperMicro server every year or so and it was keeping up.
We’re using TrueNAS Free but are changing that to TrueNAS scale pretty soon, hopefully. We have a 2x Epyc system with HHHL SSDs operating in caching mode, plus quite a bit of actual RAM cache (2x 4TB SSDs + 128 GB RAM). Those caches keep up pretty well for writing (burst up to 600 MB/s which is faster than we can write data) but less so for reading, they tend to slow, although that’s usually not a problem because we normally aren’t reading the whole datasets sequentially, but rather, read an image, process, read the next image, process, etc. If you’re reading multiple clients simultaneously (and I’d really encourage you to know your workload like the back of your hand before making decisions), you’d want to probably go mostly/all flash like @wendell said. Even high performance spinning rust is going to bottleneck. If your software has local caching (for instance, for my workload, we read the images in, extract particles, and cache those on the local machine for operations, and then eventually write out a handful of final structures/files that are much smaller), you can probably get away with spinning rust if you are at 12+ drives.
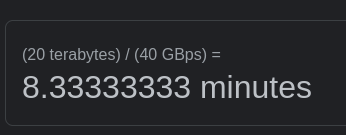
With our cameras, we are writing out a 5-10 GB image every…30 seconds or so? We average about 100 images/hr during data collections, and a weekend data collection hits 20 TB+ pretty easily. We are using Globus timers to move the data every 5-10 minutes off the camera PC to the central storage, and it’s all over 10 gigabit ethernet without much trouble. That helps alleviate the load because data collection takes longer than transfer so it keeps transfer speeds from being the bottleneck.
You’re probably going to want tapes as well. It’s just not practical for a person to manage that much data and keep it available. Depending on the type of research and who is funding it, you can look into OURRstore, as you may be eligible to use it. They have a number of options but I believe it’s just a few dollars/TB for tape backups and you just upload the files and they handle the rest for you. But it needs to be NSF/NIH related, I think.
Just some thoughts, feel free to message me if you want any more specific info. Microscopy and data management are both pretty close to my heart.
Usually there’s different things happening and recording is interspersed between those things. Moving the stage, focusing, taking trial shots, etc. One of our cameras is 96 megapixel and can write 1500 FPS at 8-bit depth, but we still only average 100 images/hr even fully automated, and those images are usually 50-100 frames. So it could write 144 GB/s but it doesn’t usually.